HMy2.CIR Cells
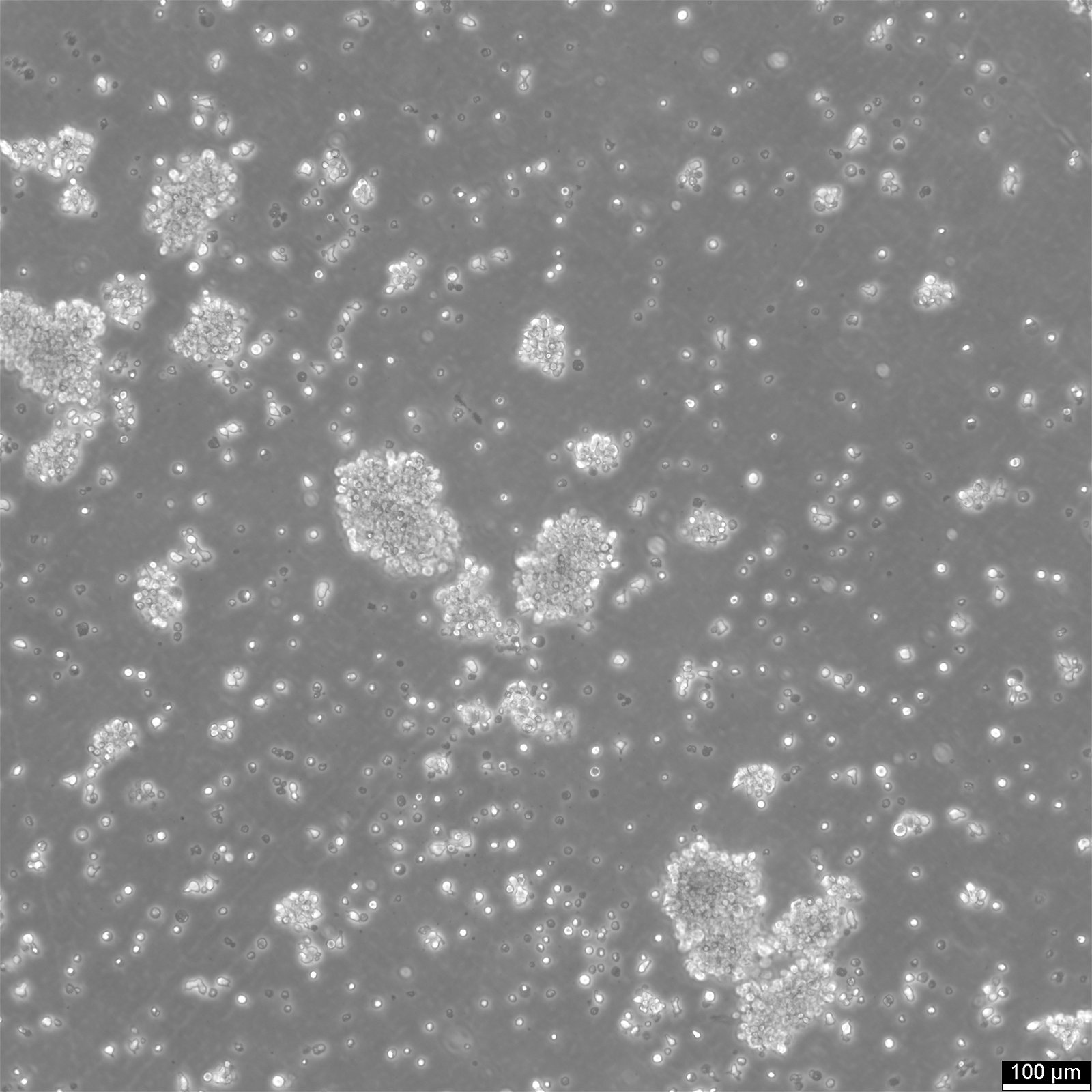

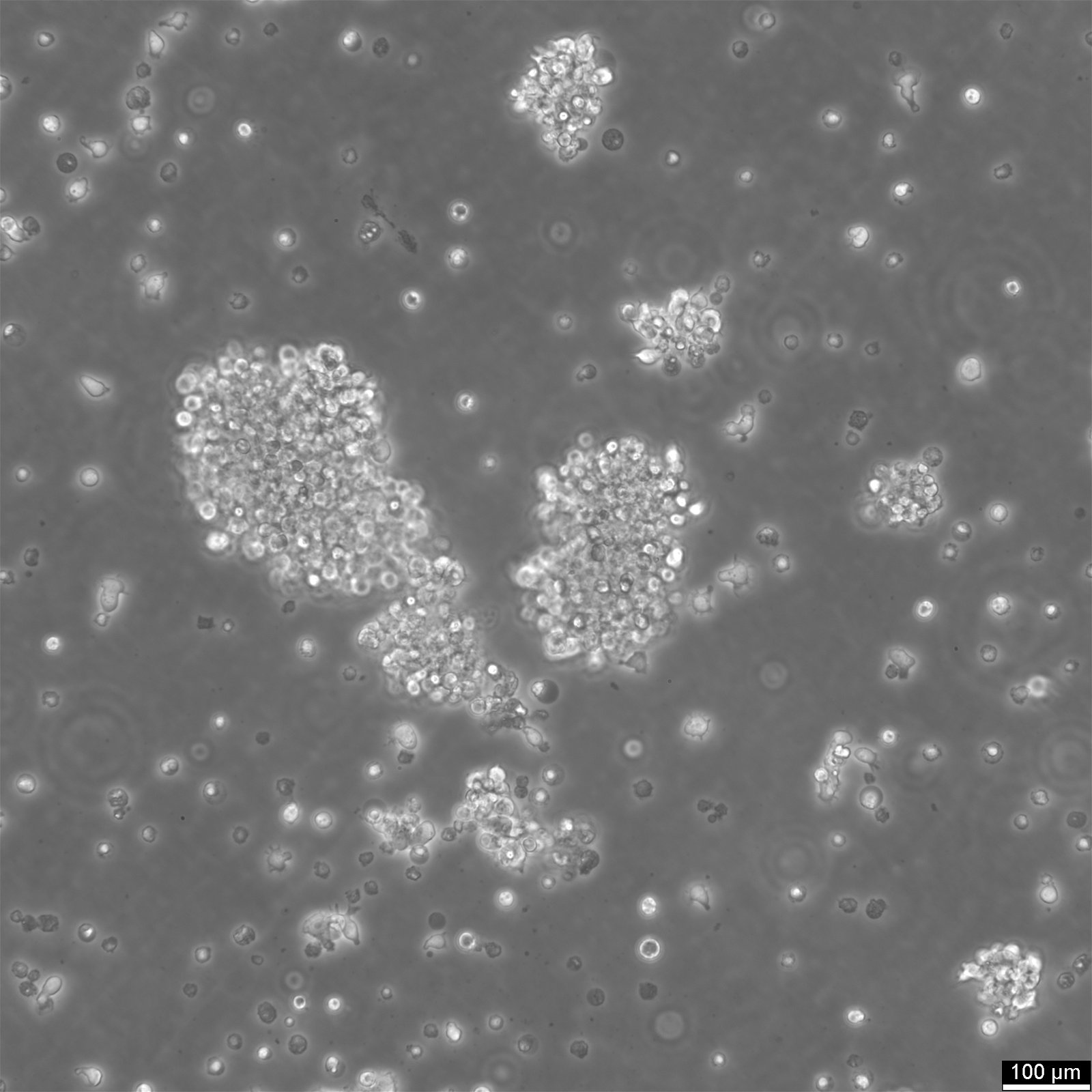






General information
| Description | This cell line is established by gamma irradiation and selection for loss of HLA class I antigen expression from the HMy.2 B lymphoblastoid cell line, a fast growing mutant of the ARH-77 cell line. The cells can be utilized as host for transfected class I major histocompatibility antigen genes. The ARH-77 cell line was reported to be positive for Epstein-Barr nuclear antigen (EBNA +) and Epstein-Barr viral capsid antigen (EBVCA +). Therefore, the HMy2.CIR cell line is assumed to be EBNA positive. The cells express small amounts of HLA Cw4 but no HLA A or B locus products. |
|---|---|
| Organism | Human |
| Tissue | B-Lymphoblast |
| Synonyms | Hmy.2 CIR, HMy2.CIR, C1R |
Characteristics
| Age | 33 years |
|---|---|
| Gender | Female |
| Ethnicity | Caucasian |
| Morphology | Lymphoblast |
| Growth properties | Suspension |
Identifiers / Biosafety / Citation
| Citation | HMy2.CIR (Cytion catalog number 305126) |
|---|---|
| Biosafety level | 2 |
Expression / Mutation
Handling
| Culture Medium | IMDM, w: 4.5 g/L Glucose, w: 4 mM L-Glutamine, w: 25 mM HEPES, w: 1.0 mM Sodium pyruvate, w: 3.024 g/L NaHCO3 (Cytion article number 820800a) |
|---|---|
| Medium supplements | Supplement the medium with 10% FBS |
| Subculturing | Gently homogenize the cell suspension in the flask by pipetting up and down, then take a representative sample to determine the cell density per ml. Dilute the suspension to achieve a cell concentration of 1 x 10^5 cells/ml with fresh culture medium, and aliquot the adjusted suspension into new flasks for further cultivation. |
| Split ratio | 1?10^5 to 1?10^6 cells/mL |
| Fluid renewal | 2 to 3 times per week |
| Freeze medium | CM-1 (Cytion catalog number 800100) or CM-ACF (Cytion catalog number 806100) |
| Handling of cryopreserved cultures |
|
Quality control / Genetic profile / HLA
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|---|
| STR profile |
Amelogenin: x,x
CSF1PO: 6,10
D13S317: 11,13
D16S539: 9,13
D5S818: 10,13
D7S820: 7,12
TH01: 8
TPOX: 8
vWA: 17
D3S1358: 16
D21S11: 29,30
D18S51: 14,16
Penta E: 12
Penta D: 10
D8S1179: 15
FGA: 20,21
D6S1043: 11,19
D2S1338: 17
D12S391: 19,20
D19S433: 14,15
|
