NCCIT Cells












General information
| Description | NCCIT was established by Shinichi Teshima (National Cancer Institute, Tokyo, Japan) in 1985 from a mediastinal mixed germ cell tumor. |
|---|---|
| Organism | Human |
| Tissue | Anterior mediastinal mixed germ cell tumor |
| Disease | Testicular embryonal carcinoma |
| Synonyms | NCC-IT |
Characteristics
| Age | 24 years |
|---|---|
| Gender | Male |
| Ethnicity | Asian |
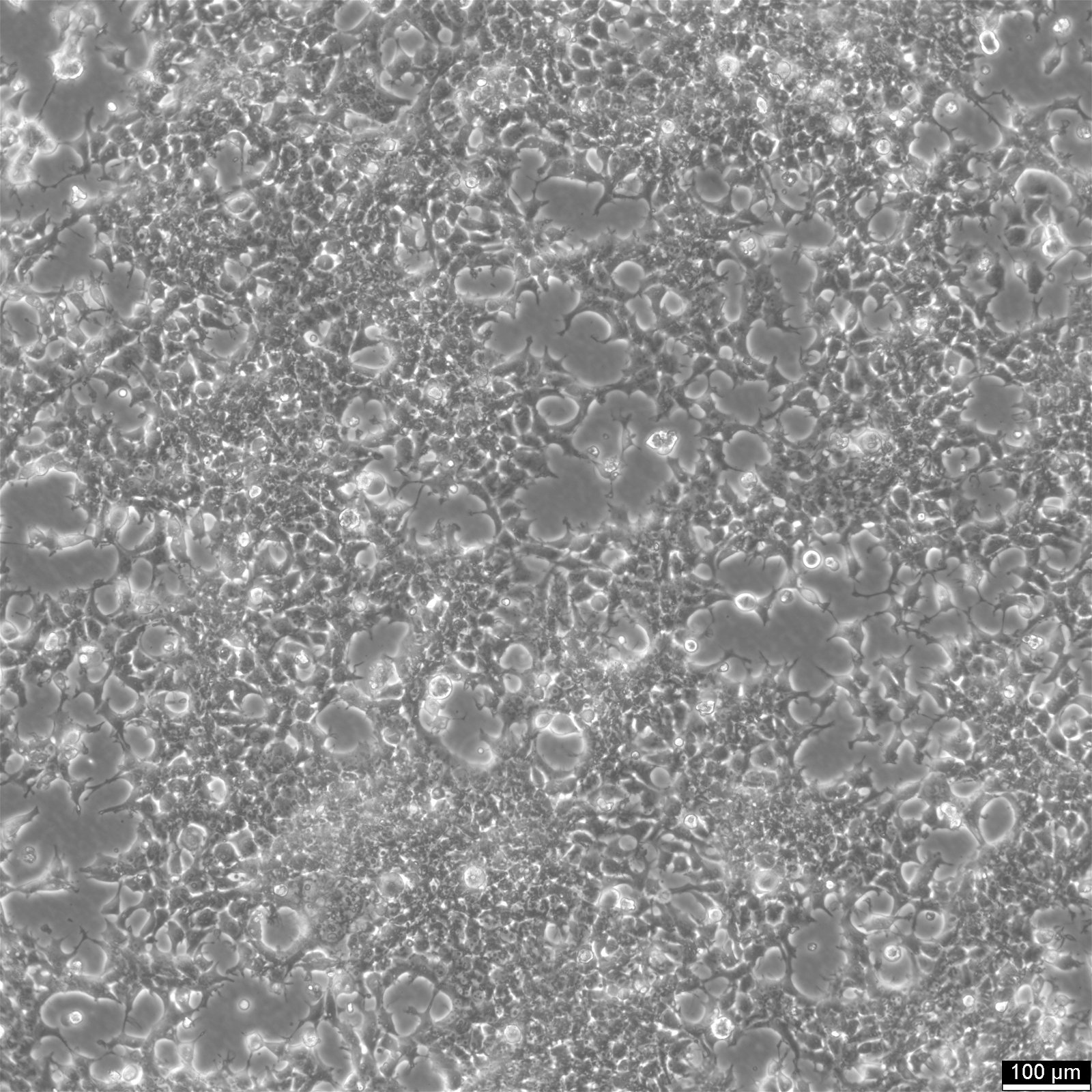
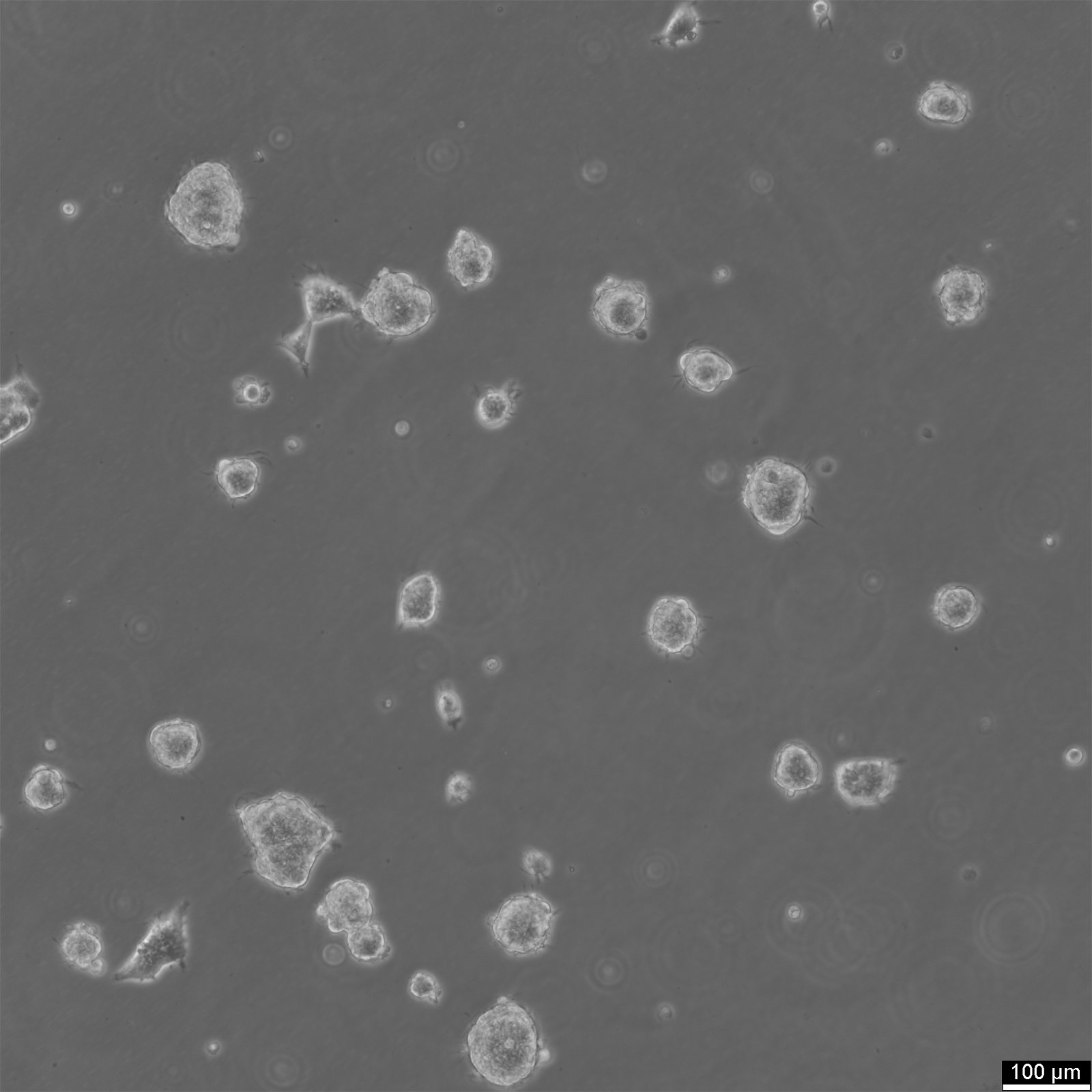
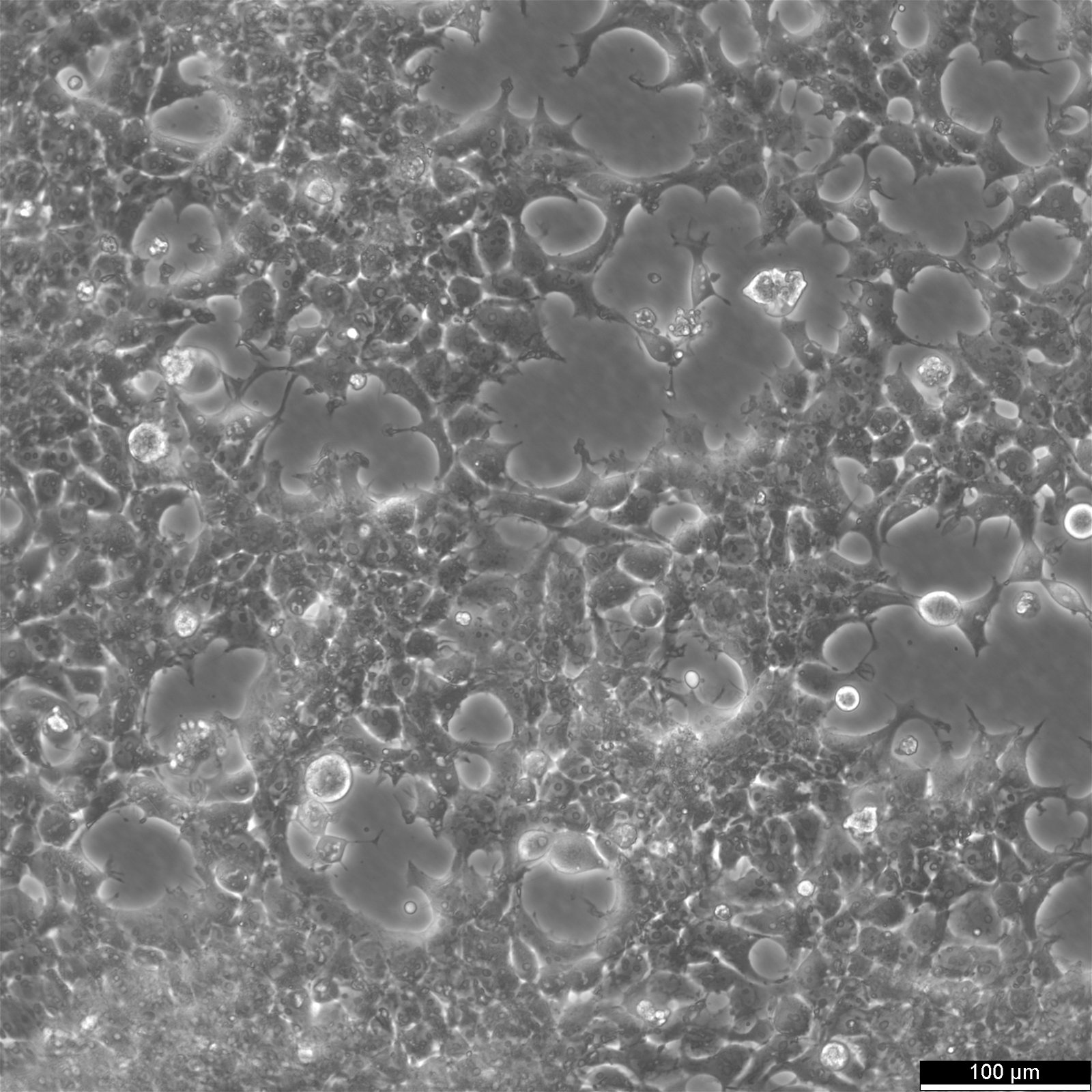
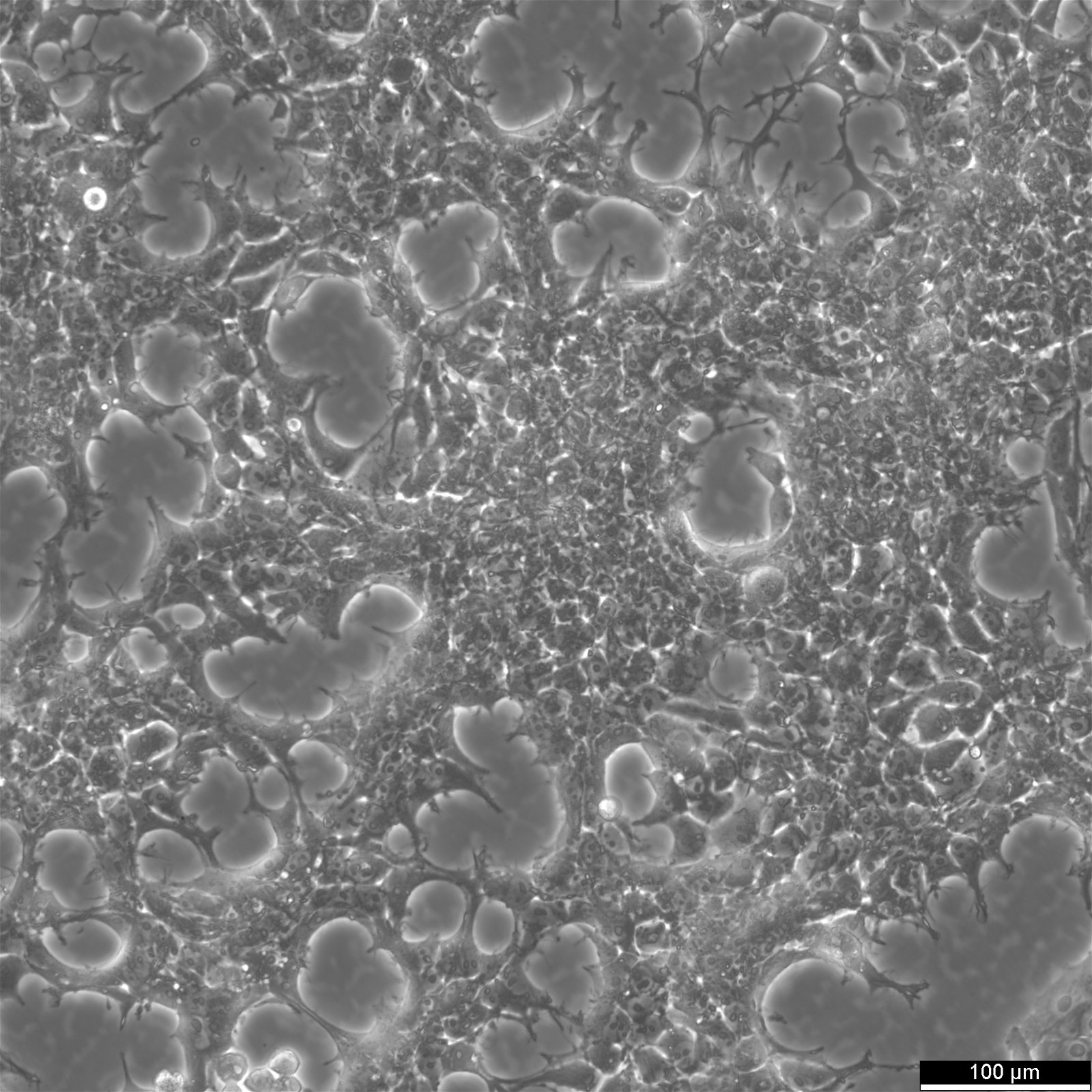
| Morphology | Epithelial |
| Growth properties | Adherent |
Identifiers / Biosafety / Citation
| Citation | NCCIT (Cytion catalog number 305080) |
|---|---|
| Biosafety level | 1 |
Expression / Mutation
Handling
| Culture Medium | RPMI 1640, w: 2.1 mM stable Glutamine, w: 2.0 g/L NaHCO3 (Cytion article number 820700a) |
|---|---|
| Medium supplements | Supplement the medium with 10% FBS |
| Passaging solution | Accutase |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Split ratio | 1:2 to 1:5 |
| Fluid renewal | 2 to 3 times per week |
| Freeze medium | CM-1 (Cytion catalog number 800100) or CM-ACF (Cytion catalog number 806100) |
| Handling of cryopreserved cultures |
|
Quality control / Genetic profile / HLA
| Sterility | Mycoplasma contamination is excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|---|
| STR profile |
Amelogenin: x,x
CSF1PO: 10,12
D13S317: 11
D16S539: 9,12
D5S818: 10,13
D7S820: 10
TH01: 7,9
TPOX: 8
vWA: 14,18
D3S1358: 16
D21S11: 29,32.2
D18S51: 13,14
Penta E: 5,14
Penta D: 10,12
D8S1179: 10,15
FGA: 22,26
D6S1043: 14
D2S1338: 18,19
D12S391: 18,22
D19S433: 13,15
|
