NCI-H460 Cells












Product number:
305020
General information
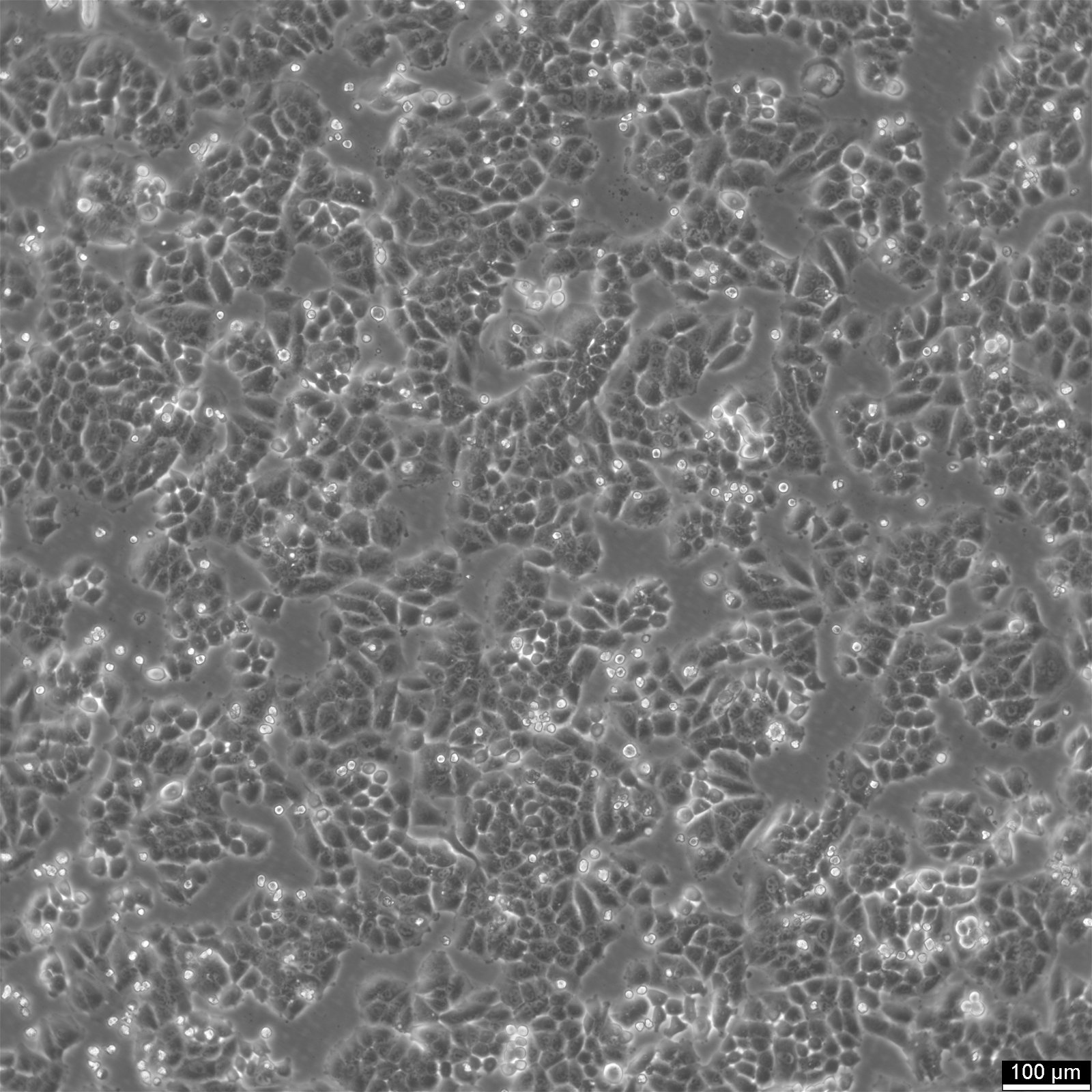
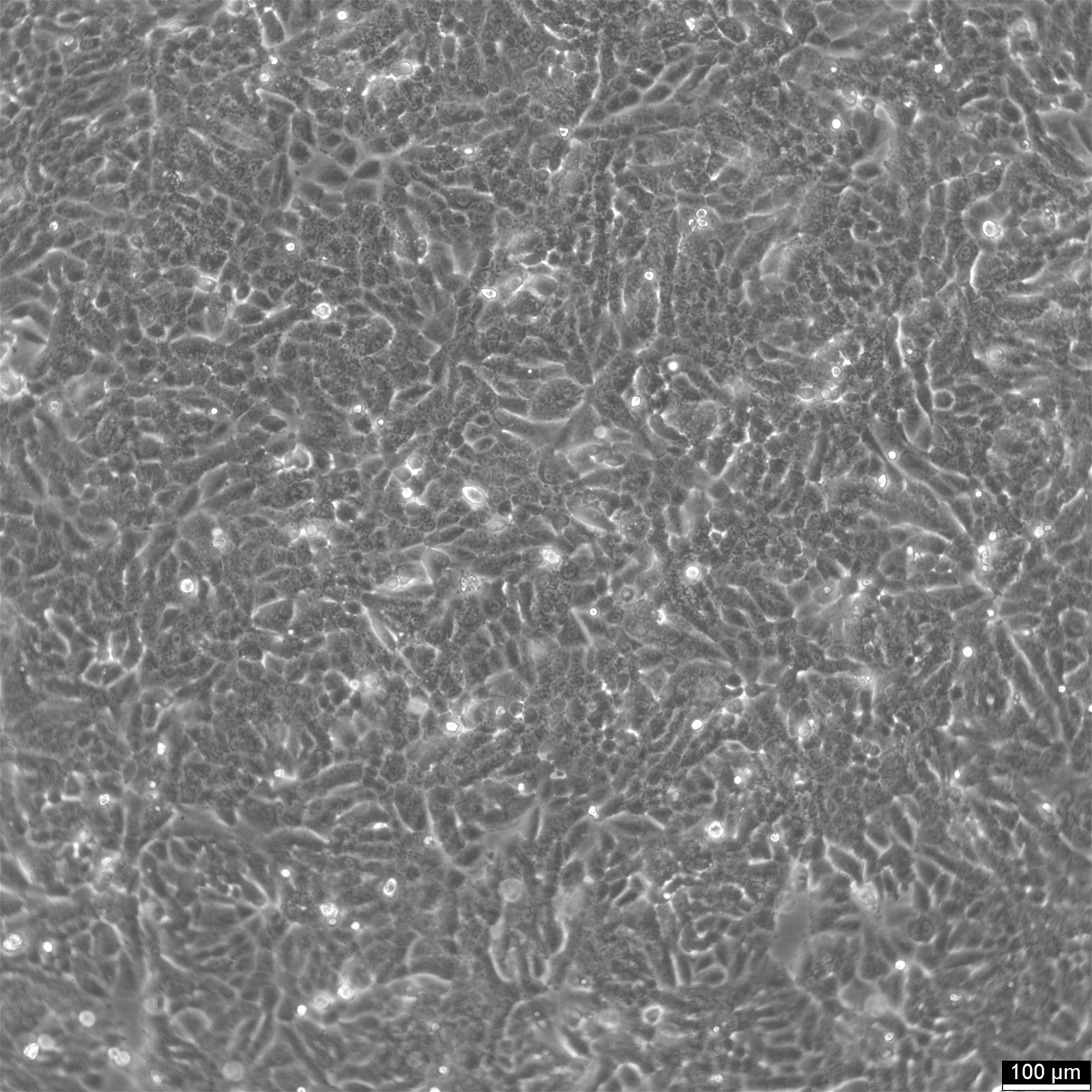
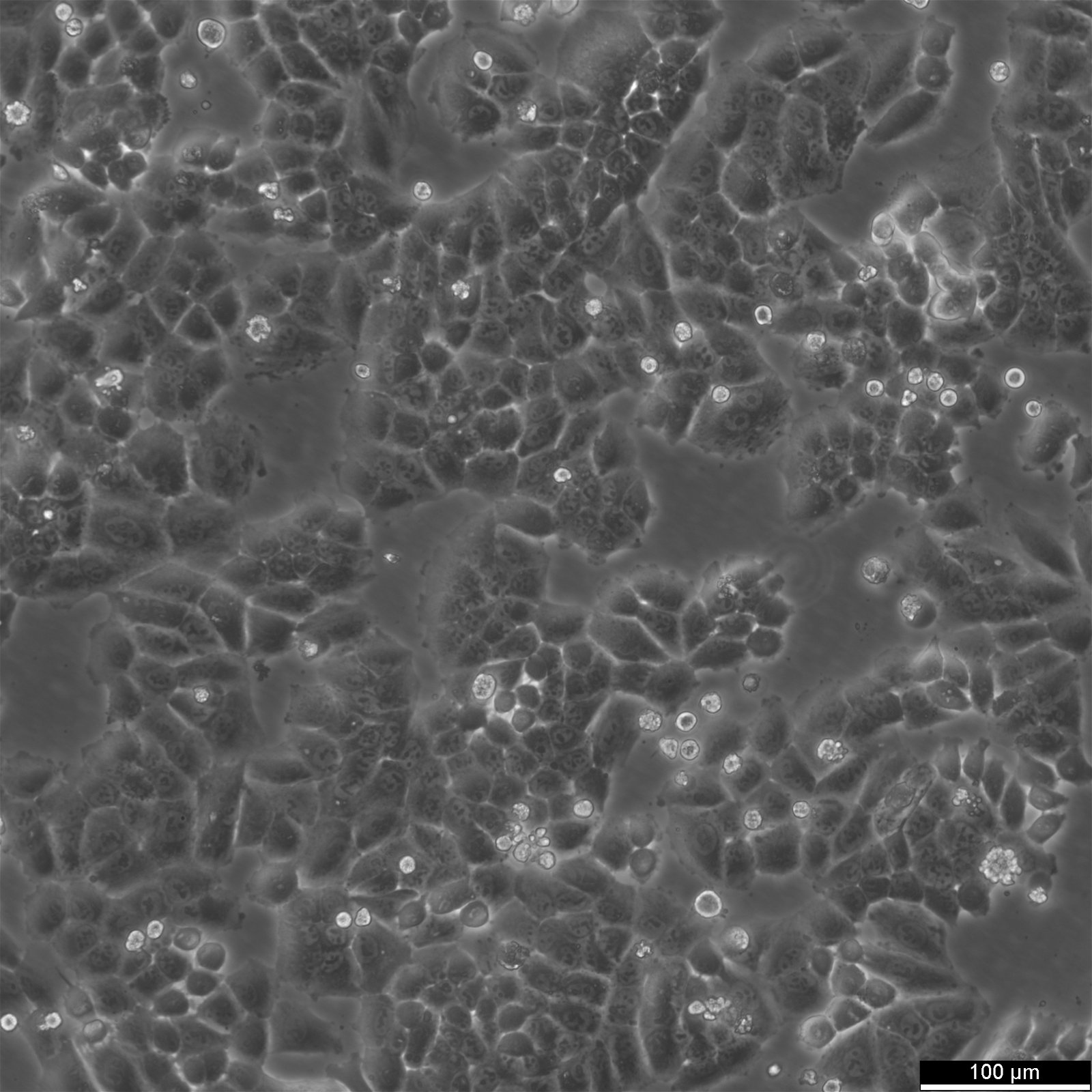
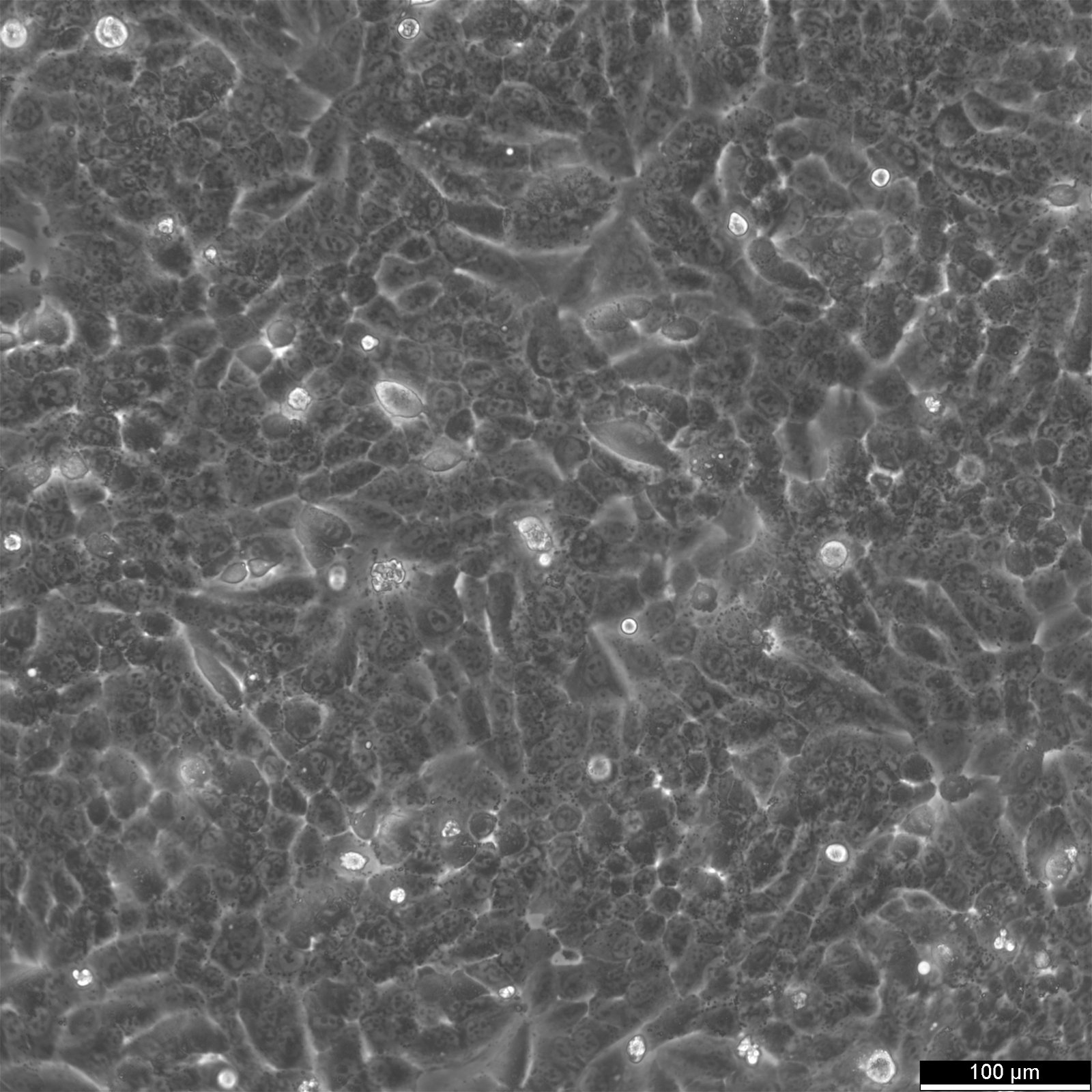
| Description | NCI-H460, also known as H460, was derived from a male patient with large cell lung carcinoma. NCI-H460 cells are adherent cells growing twice as fast than the A549 cells with a doubling time of 33 hours in RPMI 1640 supplemented with 10% FBS. They can form tumors in both in vitro and in vivo models, including nude mice. NCI-H460 cells have been shown to express p53 mRNA at high levels comparable to normal lung tissue, while exhibiting no gross structural DNA abnormalities. They stain positively for keratin and vimentin but are negative for neurofilament triplet protein. Isoenzyme analysis has shown that HPRT is localized on the surface of these non-small-cell lung cancer cell lines. AK-1, ES-D, and Me-2 isoenzymes are expressed at level 1, while G6PD and PGM1 and PGM3 isoenzymes are expressed at level B and 1-2, respectively. The cells have a hypotriploid karyotype with a modal chromosome number of 57, ranging from 53 to 65. Seven marker chromosomes are common to all cells, including der(9)t(1;9)(q21;p24), der(9)t(7;9)(p11;p22), t(10q14q), der(16)t(7;16)(q11.23;q22). Their high expression levels of p53 mRNA make them a suitable model for studying the molecular mechanisms of non-small-cell lung cancer. |
|---|---|
| Organism | Human |
| Tissue | Lung |
| Disease | Lung large cell carcinoma |
| Metastatic site | Pleural effusion |
| Synonyms | NCI-H460, NCI.H460, H-460, NCIH460, NCI-HUT-460, NCI-460 |
Characteristics
| Gender | Male |
|---|---|
| Ethnicity | European |
| Morphology | Epithelial |
| Growth properties | Adherent |
Identifiers / Biosafety / Citation
| Citation | NCI-H460 (Cytion catalog number 305020) |
|---|---|
| Biosafety level | 1 |
Expression / Mutation
| Tumorigenic | Yes |
|---|
Handling
| Culture Medium | RPMI 1640, w: 2.1 mM stable Glutamine, w: 2.0 g/L NaHCO3 (Cytion article number 820700a) |
|---|---|
| Medium supplements | Supplement the medium with 10% FBS |
| Passaging solution | Accutase |
| Subculturing | Remove the old medium from the adherent cells and wash them with PBS that lacks calcium and magnesium. For T25 flasks, use 3-5 ml of PBS, and for T75 flasks, use 5-10 ml. Then, cover the cells completely with Accutase, using 1-2 ml for T25 flasks and 2.5 ml for T75 flasks. Let the cells incubate at room temperature for 8-10 minutes to detach them. After incubation, gently mix the cells with 10 ml of medium to resuspend them, then centrifuge at 300xg for 3 minutes. Discard the supernatant, resuspend the cells in fresh medium, and transfer them into new flasks that already contain fresh medium. |
| Split ratio | 1:2 to 1:4 |
| Fluid renewal | 2 to 3 times per week |
| Freeze medium | CM-1 (Cytion catalog number 800100) or CM-ACF (Cytion catalog number 806100) |
| Handling of cryopreserved cultures | NCI-H460 cells are shipped in a deep-frozen state on dry ice. Upon receipt, confirm that the vial remains frozen. For storage, place the cryovial immediately at temperatures below -150 degrees. If you plan to culture the cells immediately, swiftly thaw the vial by shaking it in a 37 degrees water bath with clean water and an antimicrobial agent for 40-60 seconds. Remove the vial once a small ice clump persists, ensuring it remains cold. Proceed with all subsequent steps under aseptic conditions. In a sterile flow hood, disinfect the cryovial with 70% ethanol. Then, gently open the vial and transfer the cell suspension into a 15 ml centrifuge tube pre-filled with 8 ml of room temperature culture medium. Gently mix the cells. For cell separation, centrifuge at 300 x g for 3 minutes and dispose of the supernatant. Skipping centrifugation is optional, although any residual freezing medium should be removed after 24 hours. Resuspend the pellet gently in 10 ml of fresh culture medium and divide between two T25 culture flasks. Follow the subculture protocol for subsequent steps. |
Quality control / Genetic profile / HLA
| Sterility | Mycoplasma contamination is rigorously excluded using both PCR-based assays and luminescence-based mycoplasma detection methods. To ensure there is no bacterial, fungal, or yeast contamination, cell cultures are subjected to daily visual inspections. |
|---|---|
| STR profile |
Amelogenin: x,y
CSF1PO: 11,12
D13S317: 13
D16S539: 9
D5S818: 9,10
D7S820: 9,12
TH01: 9.3
TPOX: 8
vWA: 17
D3S1358: 15,18
D21S11: 30
D18S51: 13,15
Penta E: 5
Penta D: 11,13
D8S1179: 12
FGA: 21,23
D6S1043: 11,14
D2S1338: 17,25
D12S391: 21
D19S433: 14
|
